Synteny visualization and genome alignment
This tutorial walks through using JBrowse 2's dotplot and linear synteny views to visualize alignments between two genomes. These views are most useful for comparing genome assemblies, identifying large-scale rearrangements, and exploring evolutionary relationships between organisms.
The tutorial covers two main workflows: using the dotplot view for an interactive overview of chromosome-scale alignments, and using the linear synteny view for base-level inspection of aligned regions. Both views operate on PAF (Pairwise mApping Format) or similar alignment data, which you can generate with tools like minimap2 or nucmer.
For general background on synteny views and a worked example with tumor and normal genome comparison, see the SV visualization guide.
What you need
This tutorial assumes you already have a JBrowse 2 instance running (see the web quickstart for setup). You will need:
- Two genome assemblies in FASTA format (or use public assemblies)
- A whole-genome alignment in PAF format
- The jbrowse CLI to add the alignment to your config
To generate a PAF alignment yourself, install minimap2:
# Align two genomes with minimap2
minimap2 -x asm5 reference.fa query.fa > alignment.paf
The -x asm5 preset is appropriate for whole-genome assembly-to-assembly
comparison. For other comparisons (e.g., long-read to reference), use
-x map-ont, -x map-pb, or -x sr as appropriate.
Loading assemblies and alignments
Before adding an alignment, you must load both genomes as assemblies in JBrowse. If you haven't already done so, add them via the CLI:
jbrowse add-assembly reference.fa --out $OUT --load copy
jbrowse add-assembly query.fa --out $OUT --load copy
Then add the PAF alignment as a synteny track. The two assembly names passed to
-a must match the assemblies you loaded above:
jbrowse add-track alignment.paf -a query,reference --out $OUT --load copy
Important: The order of assemblies in -a query,reference matters. It
should match the order in which you ran minimap2:
minimap2 reference.fa query.fa corresponds to add-track -a query,reference.
Dotplot view
The dotplot view renders a pairwise sequence alignment as a 2D scatter plot, with one genome on each axis and aligned regions shown as diagonal lines (or off-diagonal if there are rearrangements).
Launching the dotplot
From the main JBrowse start screen, click Dotplot to open the dotplot import form. Select the two assemblies (one for each axis) and pick the synteny track to visualize.
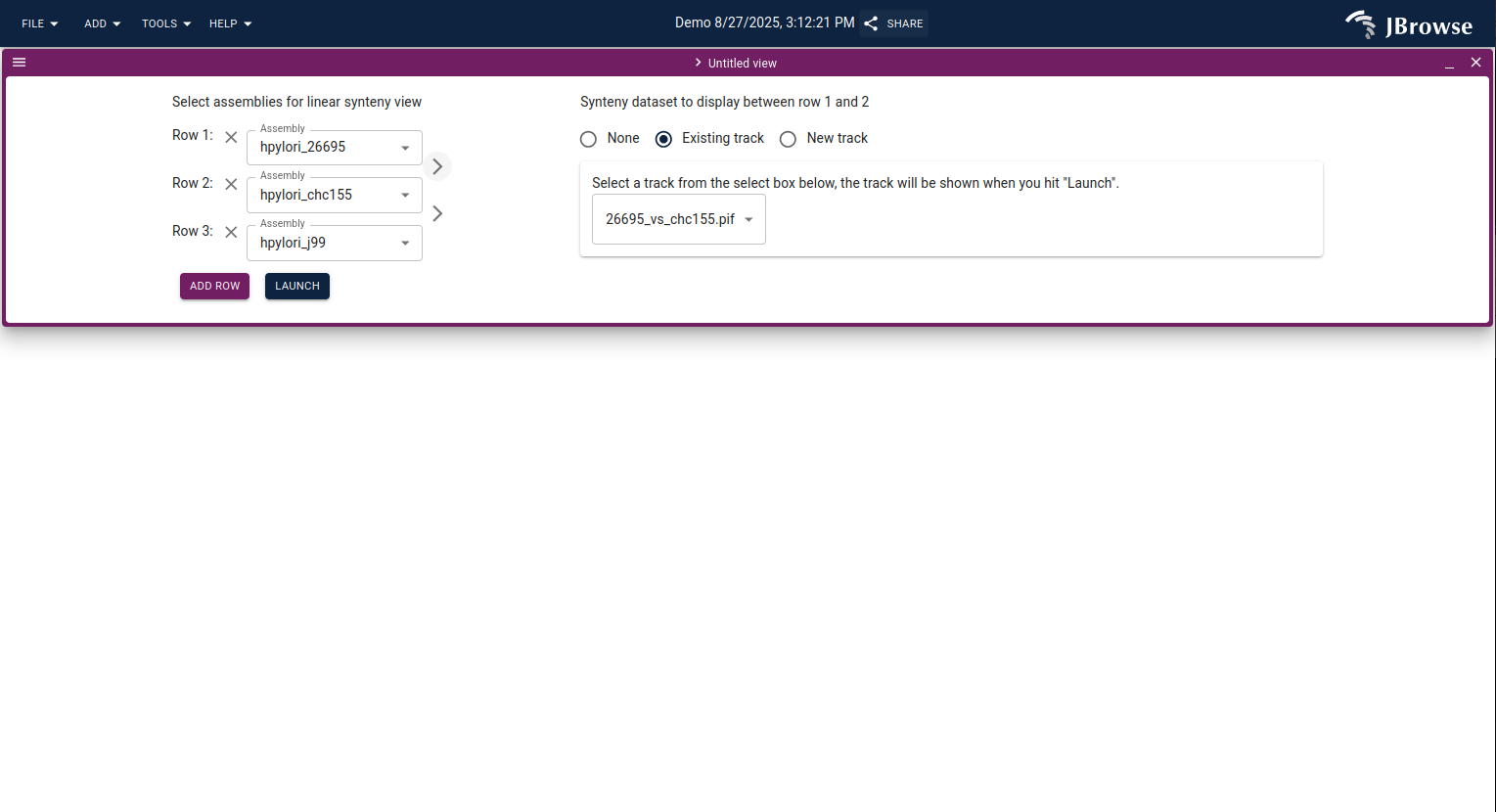
Click Open dotplot to visualize the alignment.
Reading the dotplot
In the resulting plot:
- Diagonal lines (slope ≈ 1) represent collinear aligned regions.
- Off-diagonal lines indicate rearrangements: inversions, translocations, or duplications.
- Scattered points represent short local alignments or repetitive regions.
- Gaps suggest missing or misaligned sequence.
You can zoom, pan, and click on features to get more information.
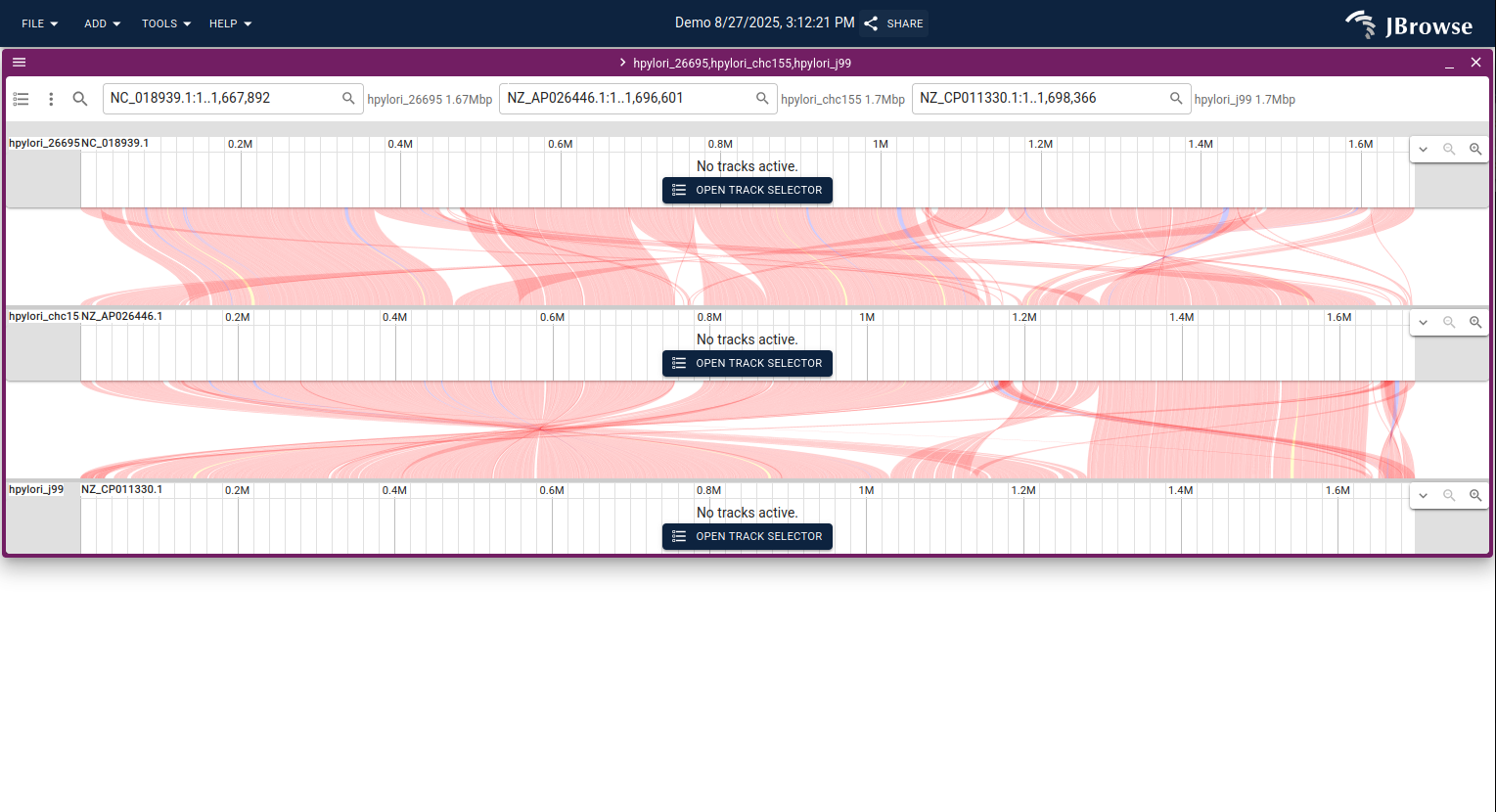
Launching a linear synteny view from the dotplot
To inspect a region in detail, click and drag over the area of interest in the dotplot, then click Launch synteny view. This opens a linear synteny view showing the base-level alignment of just that region.
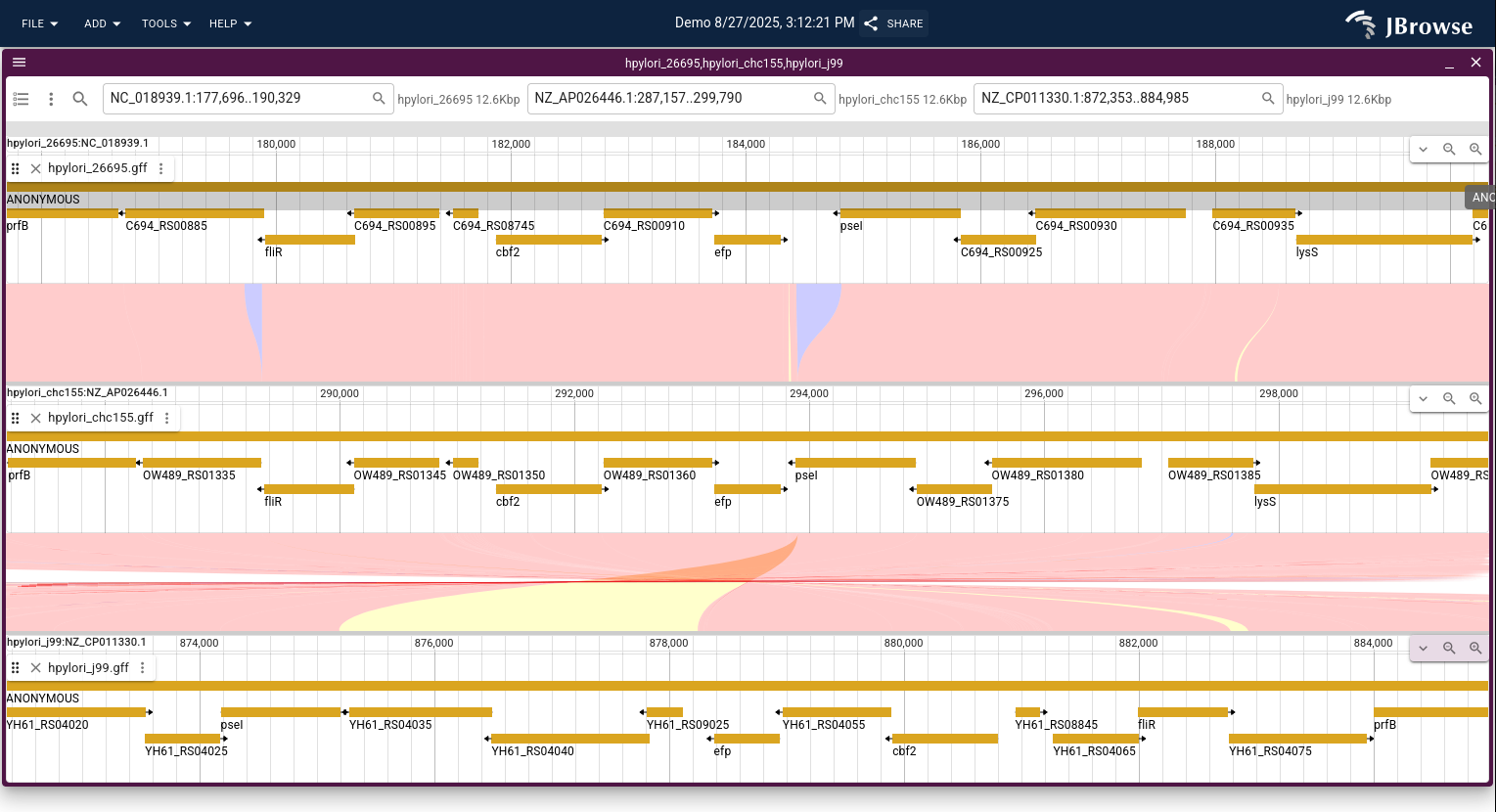
Linear synteny view
The linear synteny view shows two genomic regions aligned side-by-side, with lines connecting matching sequence blocks. This view is useful for inspecting breakpoints, identifying homologous genes, and understanding the fine structure of rearrangements.
Launching the linear synteny view
You can launch a linear synteny view in two ways:
- From the dotplot — select a region and click Launch synteny view (as shown above).
- From the start screen — click Linear synteny view, then select two assemblies and enter location ranges for each.
Configuring the linear synteny view
Once open, you can search for genomic locations in the header bar to zoom into different regions, and you can open additional tracks (e.g., gene annotations, read alignments) via the track selector.
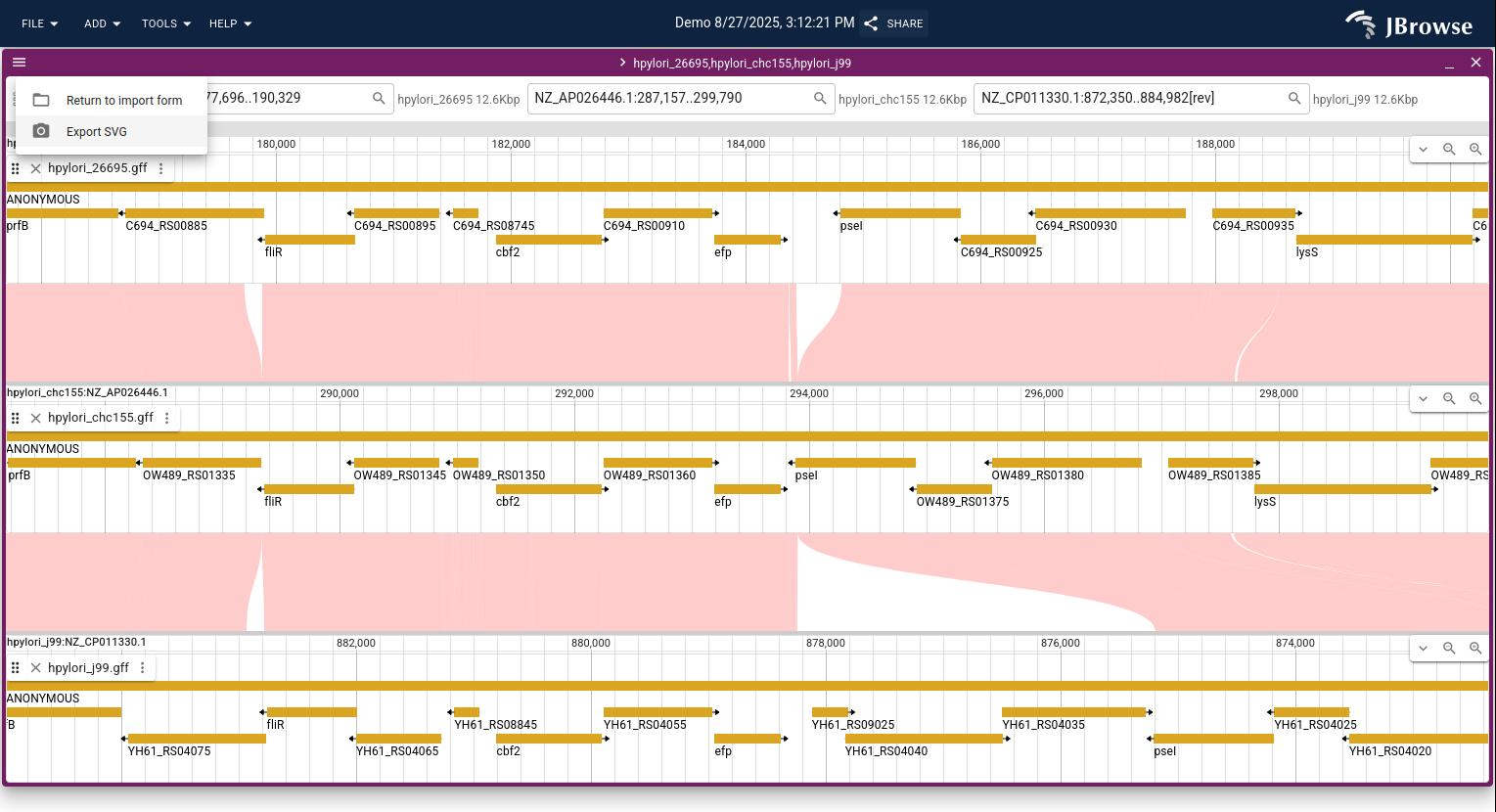
A common workflow is to open gene annotation tracks on both genomes to identify conserved syntenic genes across the alignment.
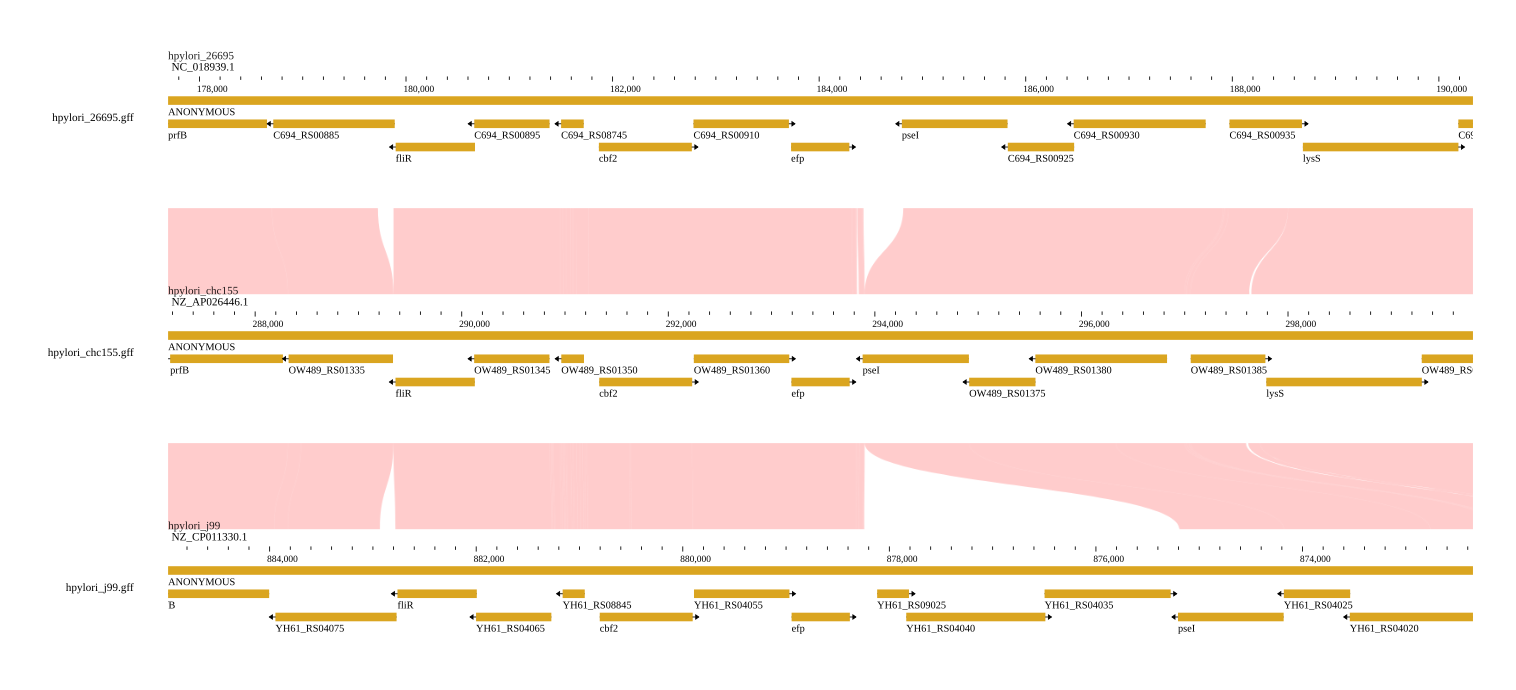
Example: Fruit crop comparison
A practical use case is comparing assemblies of related species, such as grape and peach. These organisms diverged millions of years ago but retain substantial synteny. By viewing their genomes side-by-side, you can:
- Identify conserved chromosomal regions (synteny blocks).
- Spot evolutionary rearrangements (inversions, translocations).
- Find orthologous genes between species.
The same workflows apply to other comparisons: plant strains, bacterial genomes, primate species, or tumor vs. normal genomes.
Troubleshooting
| Problem | Possible cause | Solution |
|---|---|---|
| The dotplot or synteny view is blank | Assemblies or track names don't match | Verify assembly names match your jbrowse add-assembly and add-track -a commands |
| Lines don't appear, or appear scattered randomly | The PAF was generated with wrong parameters | Try re-running minimap2 with -x asm5 for assembly comparison |
| Alignments are reversed or flipped | The PAF was generated in the opposite direction | Try swapping the order of input genomes: minimap2 query.fa reference.fa |
Next steps
- Generate your own alignments — use minimap2 with appropriate presets for your data type (assemblies, long reads, short reads).
- Overlay annotations — load gene GFF/GTF files on both sides to identify conserved genes and orthologs.
- Inspect synteny blocks — use the linear synteny view to dive into rearrangement breakpoints or conserved regions.
- Compare multiple organisms — load several pairwise alignments to build a phylogenetic perspective on genome evolution.
For more on working with synteny views, see the dotplot view guide and the linear synteny view guide.
References
Diesh, C., Stevens, G. J., Xie, P., et al. (2024). Setting Up the JBrowse 2 Genome Browser. Current Protocols, 4(8), e1120.
Diesh, C., Stevens, G. J., Xie, P., et al. (2023). JBrowse 2: A Modular Genome Browser with Views of Synteny and Structural Variation. Genome Biology, 24(1), 74.